INTRODUCTION
Over the past decades, the field of machine learning (ML) has made great strides in medicine. The greater availability of large datasetsŌĆösupported by lower data storage fees and the advent of cloud computingŌĆöhas provided a rich source of information that can be mined for ML algorithms [1]. Furthermore, enhanced computing power has accelerated the development of ML algorithms that can process complex, heterogeneous datasets involving imaging data, electronic health records, and waveforms [2]. The dramatic evolution of ML techniques has inspired researchers to build numerous prototype models for prediction, diagnosis, and prognostication. A number of these models performed equal or better in prediction and diagnosis than existing conventional statistics-based solutions. Various ML models, for example, predicted critical care outcomesŌĆöe.g., emergency department (ED) to intensive care unit (ICU) transfer and in-hospital mortalityŌĆömore accurately than existing screening tools, such as the Modified Early Warning Score, the National Early Warning Score, and the Sequential Organ Failure Assessment [3,4]. In radiology, ML-based radiomics models performed better than radiologists, especially in detecting subtle changes indescribable to the naked eye [5ŌĆō7].
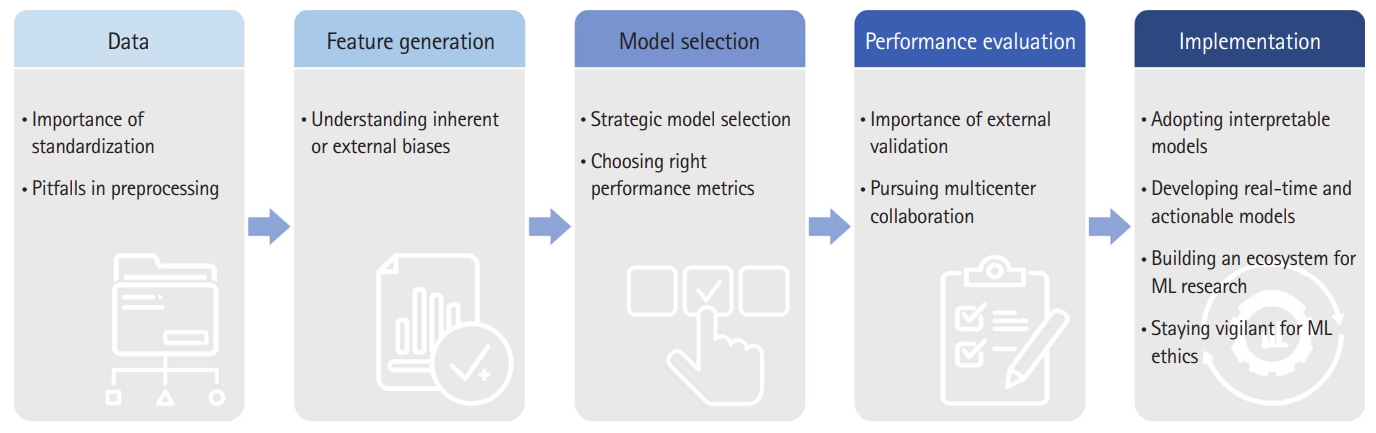
Nonetheless, several challenges must be overcome before ML algorithms can be adapted to the clinical workflow of the ICU or ED (Fig. 1). In this review, we outline these challengesŌĆöboth in developing and applying models for critical care medicineŌĆöand offer potential solutions.
CRITICAL CARE DATA
The extensive and granular datasets available in critical care medicine are promising resources for developing ML models. It is especially true when the data contain a lot of noise from the environment, such as raw vital sign data acquired from the ED. However, several challenges remain in data standardization and preprocessing.
Building a reliable ML model requires a highly structured, large, and multicenter dataset that allows for proper model training as well as internal and external validation. But obtaining such a dataset is no easy task. Electronic health records contain information collected as part of the workflow and are thus fraught with errors such as mislabeling and omission, and variation in intrahospital and interhospital reporting of clinical data creates additional challenges in data mining.
Thankfully, a number of efforts are underway to standardize data formatting. For example, Fast Healthcare Interoperability Resources (FHIR) is a preformatted healthcare database that analytics platforms can easily access and deconstruct [8]. More recently, the Critical Care Data Exchange Format (CCDEF) was developed to facilitate encoding, storing, and exchanging clinical and physiologic data across institutions globally [9]. CCDEF generates a diverse and well-represented dataset that is ideal for developing robust ML algorithms.
Once data has been gathered, optimizing the dataset through preprocessing is necessary before being employed in an ML model. Preprocessing can involve data cleaning, normalization, feature extraction, and selection to address issues with erroneous, missing, or imprecise data. Proper data preprocessing requires tremendous resources and time due to considerable size of datasets containing physiological and imaging data and directly influences the performance of the ML algorithm [10,11]. Nonetheless, the omission of preprocessing in ML studies appears common and impairs a fair assessment of the model. Even with proper preprocessing, features that may be difficult to capture in data, such as heart rate variability and interhospital differences in ICU resources, can still confound the model performance [12,13].
FEATURIZATION AND MODEL SELECTION
Featurization involves converting raw variables into numerical vectors that ML algorithms can process. Feature selection and extraction are important steps to identify clinically salient features and improve model predictability [14,15]. Therefore, features need to reflect underlying pathophysiologic mechanisms or core characteristics of data structure. For feature extraction, one needs to understand inherent biases in feature extraction (measurement error, self-reporting, human judgment) which can lead to the problem of fairness in ML. When choosing the models, they need to be built on the same well-understood variables to fairly compare their performance. Likewise, careful contextual consideration needs to be given when choosing the ML model. Another potential problem is using various ML models without consideration of data structure and study objectives, including simultaneous use of supervised and unsupervised learning just to see better performance for publication.
EVALUATION OF ML MODELS
Once the features are decided, models learn from the selected features within the different hyperparameter settings to predict desired outcomes. Choosing the best model requires calibration, which estimates concordance between the predicted probabilities and observed outcomes. Model calibration is a necessary step for measuring the relative performance of models and can assess underfitting or overfitting [16]. To evaluate fair model selection, future studies should use appropriate methods of model calibration, accounting for population size and the type of model [17]. Finally, the evaluation of a modelŌĆÖs predictive performance should assess its clinical applicability. Most of the existing studies have used the area under the receiver operating characteristics (AUROC) curve to evaluate model performance. The AUROC curve is plotted by calculating sensitivity and specificity at different thresholds. An AUROC curve provides a single performance value that is easy to interpret and compare [18]. Because the AUROC curve accounts for the true positive rate and the false positive rate, it is useful in a balanced dataset that values both positive and negative outcomes equally. The existing ML studies use the AUROC curve indiscriminately. However, the datasets used to build ML models in medicine tend to have smaller positive classes compared to negative classes. In such imbalanced datasets, the area under the precision recall curve (AUPRC) is more appropriate. The AUPRC represents positive predictive values for each true positive rate and thus focuses on positive values and is not influenced by true negatives [19]. Therefore, the use of AUPRC to evaluate models used for problems such as diagnosis, screening, and predicting mortality will lead to better estimation of the modelsŌĆÖ performance in real clinical settings.
MODEL VALIDATION
Although ML models are being rapidly developed for potential use in critical care medicine, their clinical utility is still unclear with a lack of generalizability. Small datasets, especially those produced from a single institution could lead to overfitting in a similar environment, but often not performing in other datasets. To properly implement ML algorithms in the clinical workflow of the ICU or ED, the algorithms must be externally validated. However, a recent study assessing the clinical readiness of existing ML models revealed that only 5% of the models have been externally validated [20]. Ongoing data-standardization initiatives, such as CCDEF, will hopefully integrate large datasets across multiple centers, which in turn can be employed for model validation. Further, successful model performance on prospectively collected data can demonstrate the value of ML support in clinical settings and assure clinicians of its safety.
Lack of external validation and small datasets can lead to overfitting and reduce generalizability. While curating multicenter databases in a centralized center can resolve such issues, it invites other challenges, especially in international configurations due to concerns over privacy, technical process, and data ownership. Federated learning (FL) provides a more efficient solution by allowing multiple collaborators to train models in parallel and send the updates to a central server to be integrated into a consensus model [21]. Recently, 20 centers across the globe collaborated to build a comprehensive FL model for predicting outcomes from COVID-19 infection [22]. Trained on electronic medical records and chest x-ray images, the FL model performed 16% better than a locally-trained model in predicting 24-hour oxygen treatment, with improved generalizability of 25.3%. As above, data sharing in a federated environment could overcome the limitation of external validation where data governance and privacy become obstacles.
MODEL IMPLEMENTATION
Despite a great deal of evolution in ML, several challenges still remain before their deployment. When the models are deployed at the bedside to alert physicians of impending crises, for example, their overt sensitivity can cause unintended harm. Excessive alarms that do not require cliniciansŌĆÖ immediate awareness can lead to missed real events. The ML-based alerting tools should be designed in a judicious manner to maximize the accuracy of the alarms.
Although not required, model interpretability could have a paramount impact on successful implementation. To implement a model in a clinical setting, its decisions must be verified by clinicians before use. An earlier study tested ML algorithms to build a model that could triage patients with pneumonia and predict mortality. Among the algorithms evaluated in the study, multitask neural networks were deemed to be the most accurate [23]. However, later analysis revealed a pattern in the algorithm that linked asthma to lower mortalityŌĆöexplained by the fact that patients with asthma received more attentive care and close monitoring, thus leading to better outcomes.
Since the study was published, efforts have been made to improve the interpretability of ML models. Prescience, a complex ML algorithm, accurately predicts the near-term risk of hypoxemia during surgery and displays the specific risk factors that informed its prediction [24]. The model is built using a gradient boosting machine based on both static featuresŌĆösuch as body mass index, age, and sexŌĆöand dynamic parametric values, such as tidal volume and vital signs. The impact of each feature is assigned Sharply values, which makes the predictions more interpretable through the concept used in the game theory.
The successful implementation of ML models relies on cliniciansŌĆÖ confidence in the models, which depends on how well users can explain the modelsŌĆÖ decision-making process [25]. For an ML model to play a supportive role to physicians, it is paramount to focus on features that are available real time and actionable in clinical settings. Therefore, researchers and developers should involve clinicians in an early phase of design to facilitate smooth integration into clinical workflow [26,27].
As seen from the above examples, the ML model implementation in ICU or ED population still could be far-fetched from the practice pattern of clinicians. To address that aspect from the end-user standpoint, the US Food and Drug Administration (FDA) under the Department of Health and Human Services has published an action plan for the use of artificial intelligence and ML, specifically in the form of ŌĆ£Software as a Medical Device (SaMD).ŌĆØ In the white paper, the FDA argued the need for the ŌĆ£Predetermined Change Control PlanŌĆØ to assure the quality of usable ML models for patient care [28]. Similar efforts could be seen in the Bridge2AI, a US National Institutes of Health funding initiative to promote the ML modeling environment at a multicenter level [29]. Filling the gap between the developerŌĆÖs machine to the real world remains to be a huge challenge for healthcare researchers.
Lastly, For the ML model to be successfully implemented at the bedside and performed in the realm of current clinical practice, not only the ML researchers but also clinicians (end-users) need to understand the ethical aspects of adopting it. First, all data and system-related biases should be minimized with vigilance. Bias could include a thorough examination of the environment where the model was initially developed, identification of inadequate perpetuation of systematic errors abundant in different types of healthcare practices, and so on [30]. Secondly, both researchers and clinicians need to recognize that patients and colleagues might not accept or adhere to the results of the ML model. In a survey, around 60% of Americans feel uncomfortable using artificial intelligence-driven healthcare and are also suspicious that the ML model could improve their outcomes [31]. More studies are required to understand the underlying characteristics of barriers to accepting ML in practice. Thirdly, clinicians still should strive for excellence in patient care in their traditional ways. This due diligence is mainly to avoid the moral hazards smoldering when high-performing ML models become functional at the bedside.
CONCLUSION
Research in the application of ML to critical care or emergency medicine has seen tremendous growth owing to the increasing availability of highly granular, large critical care databases. Nevertheless, for the ML-based models to serve as reliable decision support tools, the steps involved in model building to final implementation must be carefully examined. As the validity of the models is entirely dependent on the datasets, standardization of the data-gathering process and proper preprocessing of the datasets are imperative. A large portion of published studies lack a description of the preprocessing and featurization bringing into question the clinical saliency of the features involved in model performance. Moreover, selecting the right model should involve model calibration to objectively compare the accuracy of the model prediction. Even after choosing the well-calibrated model, many of the earlier studies failed to have the models externally validated which may result in the models overfitting the testing dataset reducing generalizability. In addition, the decision-making process of the algorithms should be explainable. Lastly, the endeavor to create a sustainable and scalable ecosystem should be pursued across different healthcare systems where ethical datasets could be collected and shared for fair ML research.